Standardization of nuclear run-on assay (a) 32P UTP incorporated total... | Download Scientific Diagram

Analysis of Viral Promoter Usage in EBV-Infected Cell Lines: A Comparison of qPCR Following Conventional RNA Isolation and Nuclear Run-On Assay | SpringerLink

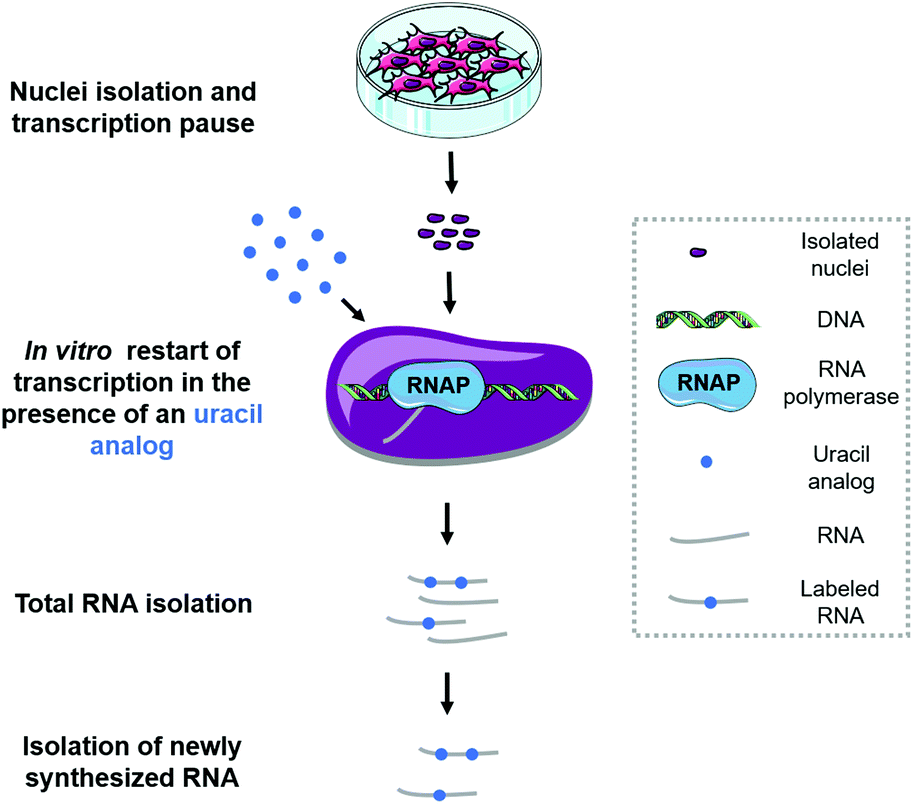
Quantification of nascent transcription by bromouridine immunocapture nuclear run-on RT-qPCR | Nature Protocols

Gene-specific quantification of nascent transcription following targeted degradation of endogenous proteins in cultured cells - ScienceDirect

Identifying Transcription Error-Enriched Genomic Loci Using Nuclear Run-on Circular-Sequencing Coupled with Background Error Modeling - ScienceDirect
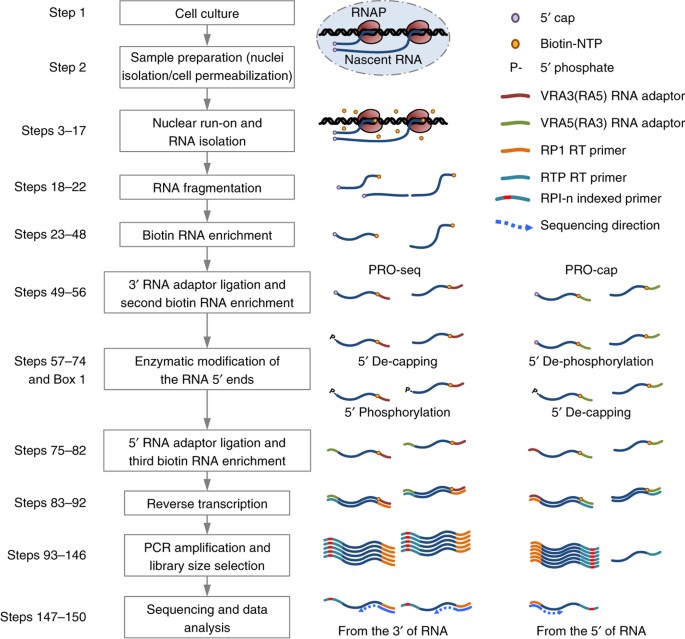
Base-pair-resolution genome-wide mapping of active RNA polymerases using precision nuclear run-on (PRO-seq) | Nature Protocols

Nuclear Run-On Assays Performed with Different UV Doses Dot blots of... | Download Scientific Diagram

Identifying Transcription Error-Enriched Genomic Loci Using Nuclear Run-on Circular-Sequencing Coupled with Background Error Modeling - ScienceDirect